ISSN: 0970-938X (Print) | 0976-1683 (Electronic)
Biomedical Research
An International Journal of Medical Sciences
Research Article - Biomedical Research (2017) Volume 28, Issue 11
Isolation and genome sequence analysis of a bacterium degrading dexamethasone
1Department of Pathogenic Biology, School of Basic Medical Sciences, Chongqing Medical University, Chongqing, PR China
2Laboratory of Pathogenic Biology and Immunology, Center of Experimental Teaching, School of Basic Medical Sciences, Chongqing Medical University, Chongqing, PR China
3Center of Pharmacy Experiment, Chongqing Medical University, Chongqing, PR China
- *Corresponding Author:
- Zhibang Yang
Department of Pathogenic Biology
School of Basic Medical Sciences, Chongqing Medical University, PR China
Accepted date: March 23, 2017
In this study, we reported the genome sequence of Burkholderia sp.CQ001, a Dexamethasone (DXM) degrading bacterium isolated from hospital wastewater and identified by morphological analysis, staining and 16S rRNA sequencing. The degradation rates of dexamethasone sodium phosphate and dexamethasone were 84.8% and 77.11% respectively. And degradation peak appeared at 24 h during agitation culture at 37°C in medium with an initial pH value of 7.5. Genome sequencing was performed using Illumina Hiseq4000 high-throughput sequencing platform. Genome sequencing results demonstrated that the bacteria had a total genome size of 7570308 bp, 66.9% G+C content. Metabolic related genes account for 80.3%. Metabolism-associated genes covered 116 metabolic pathways, including metabolic pathways of microorganisms in different environments and decomposition pathways of secondary metabolites. One of the eight key genes annotated to the metabolic pathway of steroid compounds. This is the first report on Burkholderia sp.CQ001 showing characteristics of degrading dexamethasone. Our findings may provide insights on dexamethasone degradation mechanisms, and facilitate the establishment of bioremediation engineered bacteria to eliminate the dexamethasone pollution.
Keywords
Dexamethasone, Steroid hormones, Degradation, Burkholderia, Genome sequence.
Introduction
Dexamethasone (DXM), synthetic long-acting glucocorticoids, has been extensively used in the prevention and treatment of various human and poultry diseases in the past decades [1-3]. Dexamethasone Sodium Phosphate (DSP) is the most commonly used formulation of DXM in clinical practice due to its water solubility and absorbability [4]. However, long-term or high-dose use of DXM frequently leads to severe adverse events including Cushing's syndrome, cardiovascular disease, osteoporosis, or aseptic necrosis of the femoral head [5-7]. Previous studies have reported that DXM residues may cause environmental pollution through a variety of ways including industrial, hospital, or domestic wastewater, or even drinking water [8-11]. Therefore, studies on dexamethasone degradation mechanisms are important in establishment of bioremediation engineered bacteria to eliminate the dexamethasone pollution.
Burkholderia is a gram-negative bacterium distributed widely in water, soil, plants and human body [12,13]. It has a unique metabolic potential that can grow and reproduce with more than 200 kinds of organic compounds as a carbon source [13]. The genome sequences of some strains of the Burkholderia genus have been disclosed, and the whole genomic information of B. vietnamiensis strain G4 and B. xenovorans LB400 with the property of degradation have been reported [14]. However, the degrading effects on steroids and their related genes by these strains have not been investigated.
In this study, a strain using dexamethasone sodium phosphate as the sole carbon source and energy sources were successful obtained and identified as Burkholderia and was named as Burkholderia sp.CQ001. Based on the genome sequence and bioinformatics analysis, we found that Burkholderia sp.CQ001 had genes and metabolic pathways that can potentially degrade dexamethasone steroids with high efficiency.
Experiments
Isolation and identification of bacteria
One bacterial strain was isolated from the wastewater of a hospital in Chongqing, China (29° 33'N, 106° 28'E) using the methods reported by Wang et al. [15]. The solid phase extraction-High Performance Liquid Chromatography (HPLC) method was used to detect the degradation effects of the bacteria on DXM and DSP (500 μg/ml), and the effects of initial medium pH and temperature on the degradation.
Finally, the morphological features of bacteria were observed by electron microscopy and scanning electron microscopy after Gram-staining. The main biochemical reactions were detected by conventional methods. The taxonomic status of the bacteria was identified by 16s rRNA sequencing.
Homology and phylogenetic analysis
Genomic DNA was extracted using DNA preparation kit according to the manufacturer’s instructions (Takara, Japan). The 16s rRNA gene sequences of Burkholderia sp.CQ001 were amplified by PCR with universal primers of 27F and 1492R. The genomic sequencing for amplification products was conducted using sequencer of ABI3730XL at Majorbio Biopharm Technology Company (Shanghai, China), and sequence analysis was carried out in the NCBI database. Meanwhile, The 16s rRNA full sequences of 16 bacterial strains with high similarity to the sequenced strains were downloaded from NCBI GenBank, and the multiple sequence comparison was conducted using CLUSTAL W [16]. Ultimately, comparative results were analysed by MEGA 6.0 [17] using neighborjoining method (bootstrap=1000) to build the phylogenetic tree [18].
Genome sequencing and assembly
Genome sequencing was performed at Majorbio Bio-Pharm Technology company (Shanghai, China) using Illumina Hiseq4000 high-throughput sequencing platform by building a ~500 bp Paired-End library. After handling of the raw data, the optimized sequences were spliced using SOAPdenovo software (version 2.04) [19]. The partial filling and base correction were carried out for the optimal assembly sequences obtained through the above-mentioned procedures by GapCloser software (version 1.12). Ultimately, the filled and corrected sequences were matched to the measured sequences (reads), and assembled results were evaluated by statistics analysis for the depth of coverage between the content of G+C and reads. A total of 1,390,912 of original sequences were identified, and 12,891,489 high-quality sequences were obtained after quality cutting.
Genome annotation
The rRNAs and tRNAs genes included in the genome were predicted using RNAmmer (version 1.2) [20] and tRNAscan- SE (version 1.3.1) [21] software, respectively. The proteincoding sequence was predicted using Glimmer software (version 3.02) [22]. To obtain functional annotation information of COG [23], GO, KEGG [24] and other relevant database, a blast alignment (BLAST 2.2.28+) was conducted between the predicted sequences and Nr, genes, string and GO database. Blast alignment information was adopted only when E values ≤ 10-5. The relevant functional genes that may be involved in the degradation of DXM were assessed.
Comparative genomics analysis
Next, we analysed the basic genomic features of obtained sequences including gene sequence length (bp), G+C percentage, number of predicted genes, and number of encoding RNA (tRNA and rRNA). Then, these features of obtained sequences were compared to the sequences of 6 representative strains of Burkholderia genus selected from NCBI database.
Results and Discussion
Biological characteristics of bacteria
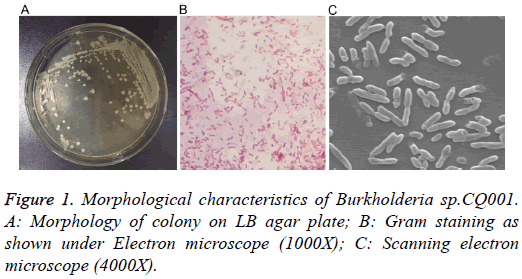
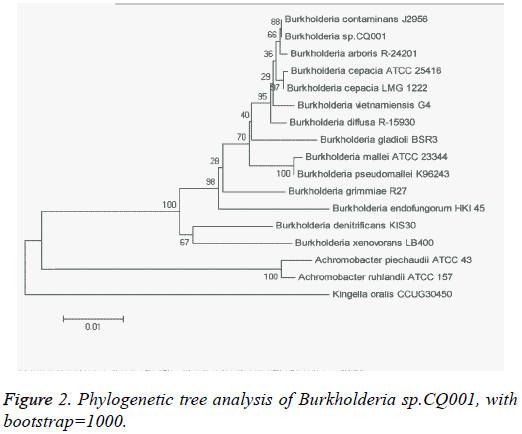
The isolated bacteria with potential of degrading DXM/DSP were gram-negative bacillus with straight or slightly curved cylinder shape, and 6-15 μm × 4-15 μm in length and diameter. On LB agar plates, the bacteria formed a milky white opaque colony with medium size (d=1.5-2.0 mm), round, smooth and moist surface, regular edge, and easy to be stirred up (Figure 1). The catalase and oxidase tests showed positive results. The bacteria were initially identified by 16s rRNA sequencing, which belonged to the genus Burkholderia and were named as Burkholderia sp.CQ001. A 16srRNA-based phylogenetic tree was constructed (Figure 2), which revealed Burkholderia sp.CQ001 having closest kinship with Burkholderia contaminants J2956 in evolution.
These bacteria can degrade DSP into DXM and then degrade the latter into other small molecules, with the degradation rates of 84.8% and 77.11%, respectively. And degradation peak appeared at 24 h during agitation culture at 37°C in medium with an initial pH value of 7.5.
The basic characteristics of the genome
Burkholderia sp.CQ001 had a total genome size of 7570308 bp, and contained 66.9% G+C. The 8,705 predicted genes included 8,632 protein-coding genes and 73 RNA coding genes (15 rRNA and 58 tRNA). The statistical information of gene prediction results was demonstrated in Table 1.
| Features | Number |
|---|---|
| Gene number | 8705 |
| Gene total length | 7570308 bp |
| Gene average length | 869 bp |
| Gene density | 0.946 genes per kb |
| GC content in gene region (%) | 66.9 |
| Gene/Geonme (%) | 82.3 |
Table 1: General features of Burkholderia sp. CQ001 genome.
Gene annotation and function classification
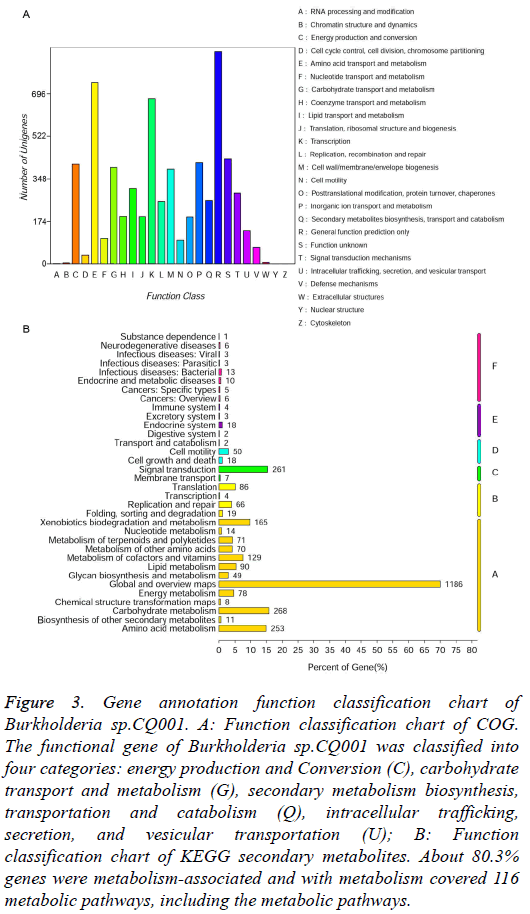
COG function classification: The alignment results compared with COG sequences were shown in Figure 3A, about 70.1% genes were annotated. There were 408 genes for energy production and Conversion (C), 395 for carbohydrate transport and metabolism (G), 260 for secondary metabolism biosynthesis, transportation and catabolism (Q), 135 for intracellular trafficking, secretion and vesicular transportation (U). These genes may be closely related to the degradation of dexamethasone of the bacteria. Depending on COG classification, the functional gene of Burkholderia sp.CQ001 was classified into four categories (Table 2). Therefore, we speculate that the bacteria have a unique metabolism potential that can grow and reproduce with DSP as the sole carbon source and survive in the extreme environment just like hospital wastewater.
| Category | Group | Function | Number of genes |
|---|---|---|---|
| Information storage and processing | A | RNA processing and modification | 1 |
| B | Chromatin structure and dynamics | 4 | |
| J | Translation, ribosomal structure and biogenesis | 193 | |
| K | Transcription | 676 | |
| L | Replication, recombination and repair | 253 | |
| Cellular processes and signaling | D | Cell cycle control, cell division, chromosome partitioning | 35 |
| M | Cell wall/membrane/envelope biogenesis | 388 | |
| N | Cell motility | 97 | |
| O | Posttranslational modification, protein turnover, chaperones | 192 | |
| T | Signal transduction | 290 | |
| U | Intracellular trafficking, secretion and vesicular transport | 135 | |
| V | Defense mechanisms | 68 | |
| W | Extracellular structures | 6 | |
| Metabolism | C | Energy production and conversion | 408 |
| E | Amino acid transport and metabolism | 741 | |
| F | Nucleotide transport and metabolism | 104 | |
| G | Carbohydrate transport and metabolism | 395 | |
| H | Coenzyme transport and metabolism | 194 | |
| I | Lipid transport and metabolism | 308 | |
| P | Inorganic ion transport and metabolism | 414 | |
| Q | Secondary metabolism biosynthesis,transport and catabolism | 260 | |
| Poorly characterized | R | General function prediction only | 896 |
| S | Function unknown | 429 | |
| Total | 6487 |
Table 2: COG distribution of Burkholderia sp.CQ001.
Figure 3: Gene annotation function classification chart of Burkholderia sp.CQ001. A: Function classification chart of COG. The functional gene of Burkholderia sp.CQ001 was classified into four categories: energy production and Conversion (C), carbohydrate transport and metabolism (G), secondary metabolism biosynthesis, transportation and catabolism (Q), intracellular trafficking, secretion, and vesicular transportation (U); B: Function classification chart of KEGG secondary metabolites. About 80.3% genes were metabolism-associated and with metabolism covered 116 metabolic pathways, including the metabolic pathways.
KEGG metabolic pathway the specific genes involved biological pathways can be achieved through KEGG analysis (Figure 3B). About 80.3% genes were metabolism-associated and they covered 116 metabolic pathways, including the metabolic pathways of microorganisms in different environments and decomposition pathways of secondary metabolites. More importantly, there were also 165 genes of degradation and metabolism for xenobiotics (A: Metabolism/ Xenobiotics biodegration and metabolism). Among them, 8 pathways involved in the metabolism of steroid compounds can encode 7 key enzymes (Table 3). According to the steroid degradation pathway (http://www.genome.jp/kegg/ pathway.html), the metabolic pathways of steroid compounds in Burkholderia sp.CQ001 KEGG pathway are as the follows: 3β-HSD continuously catalyses the 3β-OH steroid compounds dehydrogenation and isomerism reaction, facilitating the first step in the ring opening reaction of the steroid nucleus [25,26]. The interaction of KstD and KSH induces the cleavage of the B ring of the steroid nucleus [27]. KSH is composed of KshA and KshB. The Hsa family proteins catalyze the cleavage of the steroid nucleus into ATP and small molecule organic salts and coenzymes [28]. The presence of these genes and pathways provides a basis for the degradation of steroid hormones by Burkholderia sp.CQ001.
| Degrading enzymes | Definitions | ORF | Orthologous genes |
|---|---|---|---|
| HsaA | 3-hydroxy-9,10-secoandrosta-1,3,5(10)-triene-9,17-dione monooxygenase (EC:1.14.14.12) | orf00263_1 | K16047 |
| orf00460_1 | K16047 | ||
| HsaC | 3,4-dihydroxy-9,10-secoandrosta-1,3,5(10)-triene-9,17-dione 4,5-dioxygenase (EC:1.13.11.25) | orf00274_1 | K16049 |
| HsaD | 4,5:9,10-diseco-3-hydroxy-5,9,17-trioxoandrosta-1(10),2-diene-4-oate hydrolase (EC:3.7.1.17) | orf00306_1 | K16050 |
| KstD | 3-oxosteroid 1-dehydrogenase (EC:1.3.99.4) | orf00265_1 | K05898 |
| KshA | 3-ketosteroid-9-alpha-hydroxylase oxygenase subunitA (EC:1.14.13.142) | orf00291_1 | K15982 |
| KshB | 3-ketosteroid-9-alpha-hydroxylase reductase subunitB (EC:1.14.13.142) | orf00267_1 | K15983 |
| 3β-HSD | 3(or 17)beta-hydroxysteroid dehydrogenase (EC:1.1.1.51) | orf00350_1 | K05296 |
Table 3: Steroid compounds-degrading enzymes in Burkholderia sp.CQ001 genome.
Comparative genomics information
Burkholderia genus, with abundant genetic diversity, has more than 40 strains including bacteria that are pathogenic to human, animal and plant and bacteria can degrade organic matter. The basic genomic characteristics of Burkholderia sp.CQ001 were compared to the most representative strains selected from each of the Burkholderia strains (Table 4). Chain et al. [14] revealed that Burkholderia xenovorans LB400, a non-pathogenic bacterium isolated from the sewage, had enriched degradation pathways for aromatic compounds, including 11 major trichloromethyl aromatic compounds and more than 20 minor aromatic compounds, such as biphenyl, diaminophenol, and trichloro-cresol. Furthermore, the preliminary comparison demonstrated that Burkholderia sp.CQ001 had the closest number of predicted genes with Burkholderia xenovorans LB400, and a wealth of degradation pathways of aromatic compounds were also found in KEGG pathways analysis. Thus, we speculate that these two strains share a high degree of similarity in the gene structures and functions. However, the degradation of steroidal compound by Burkholderia xenovorans LB400 and by other Burkholderia strains has not been reported. More degradation related genes in Burkholderia sp.CQ001 may be disclosed in further comparative analysis of these two bacteria.
| Genome | Length (bp) | G+C (%) | Genes | rRNA genes | tRNA genes | Isolated | Signal P (%) | NCBI accession |
|---|---|---|---|---|---|---|---|---|
| Burkholderia xenovorans LB400 | 9702951 | 62.6334 | 8596 | 18 | 65 | Polluted water | 9.52 | GCF_000756045.1_ASM75604v1 |
| Burkholderia vietnamiensis G4 | 8391070 | 65.7381 | 7592 | 18 | 68 | Polluted water | 9.32 | GCF_000016205.1_ASM1620v1 |
| Burkholderia mallei ATCC 23344 | 6935527 | 68.4885 | 5506 | 10 | 56 | Human | 7.39 | GCF_000011705.1_ASM1170v1 |
| Burkholderia cenocepacia J2315 | 8055782 | 66.8993 | 7273 | 18 | 73 | Human | 11.48 | GCF_000009485.1_ASM948v1 |
| Burkholderia pseudomallei K96243 | 7247547 | 68.0587 | 5935 | 12 | 61 | Human | 10.34 | GCF_000011545.1_ASM1154v1 |
| Burkholderia gladioli BSR3 | 9052299 | 67.3975 | 7708 | 15 | 69 | Plants | 9.98 | GCF_000194745.1_ASM19474v1 |
| Burkholderia sp. CQ001 | 9192367 | 66.015 | 8705 | 15 | 58 | Polluted water | 7.42 | SRP073478 |
Table 4: Comparison of basic genome characteristics between Burkholderia sp.CQ001 and other strains of Burkholderia.
Conclusions
At present, the studies on the degradation mechanism of steroids are mainly involved in Streptococcus and Mycobacterium [29-31]. Among the Burkholderia, No related gene or metabolic pathway of the degradation of dexamethasone has been found. Here we report Burkholderia sp.CQ001, a Dexamethasone (DXM) degrading bacterium successfully isolated from hospital wastewater. The degradation rates of dexamethasone sodium phosphate and dexamethasone were 84.8% and 77.11% respectively. Our study gives a description of the genome sequence of Burkholderia sp.CQ001 and a preliminary analysis of related functional genes and metabolic pathways. More importantly, the findings provide reference for further studies on dexmethasone degradation mechanisms and establishment of bioremediation engineered bacteria to eliminate the dexamethasone pollution.
Registration number of gene sequences
The genome sequences of Burkholderia sp.CQ001 were stored in the DDBJ/EMBL/GenBank database under accession number PRJNA339502.
Acknowledgements
This work was supported by Science and Technology Project from Science and Technology Commission of Yuzhong District of Chongqing (No. 20160110).
Conflict of Interest
The authors declare no conflict of interest.
References
- Oishi Y, Fu ZW, Ohnuki Y, Kato H, Noguchi T. Molecular basis of the alteration in skin collagen metabolism in response to in vivo dexamethasone treatment: effects on the synthesis of collagen type I and III, collagenase, and tissue inhibitors of metalloproteinases. Br J Dermatol 2002; 147: 859-868.
- Ursula G, Claudia V, Francesca F, Bozza S, Bianchi R, Vacca C, Orabona C, Belladonna ML, Ayroldi E, Nocentini G, Boon L, Bistoni F, Fioretti MC, Romani L, Riccardi C, Puccetti P. Reverse signaling through GITR ligand enables dexamethasone to activate IDO in allergy. Nat Med 2007; 13: 579-586.
- Lim YJ, Jung JW. Clinical outcomes of initial dexamethasone treatment combined with a single high dose of intravenous immunoglobulin for primary treatment of Kawasaki disease. Yonsei Med J 2014; 55: 1260-1266.
- Tomida H, Yotsuyanagi T, Ikeda I. Solubilization of steroid hormones by polyoxyethylene lauryl ether. Chem Pharm Bull 2013; 26: 2832-2837.
- Thibier M, Rolland O. The effect of dexamethasone (DXM) on circulating Testosterone (T) and Luteinizing Hormone (LH) in young postpubertal bulls. Theriogenology 1976; 5: 53-60.
- Canalis E, Mazziotti G, Giustina A, Bilezikian JP. Glucocorticoid-induced osteoporosis: pathophysiology and therapy. Osteoporosis Int 2007; 18: 1319-1328.
- Duff BA, Chun KP, Ma D, Lythgoe MF, Scott RC. Dexamethasone exacerbates cerebral edema and brain injury following lithium-pilocarpine induced status epilepticus. Neurobiol Dis 2014; 63: 229-236.
- Chang H, Hu J, Shao B. Occurrence of natural and synthetic glucocorticoids in sewage treatment plants and receiving river waters. Environ SciTechnol 2007; 41: 3462-3468.
- Van der Linden SC, Heringa MB, Man HY, Sonneveld E, Puijker LM, Brouwer A, Van der Burg B. Detection of multiple hormonal activities in wastewater effluents and surface water, using a panel of steroid receptor CALUX bioassays. Environ SciTechnol 2008; 42: 5814-5820.
- Schriks M, van Leerdam JA, van der Linden SC, van der Burg B, van Wezel AP, de Voogt P. High-resolution mass spectrometric identification and quantification of glucocorticoid compounds in various wastewaters in the Netherlands. Environ SciTechnol 2010; 44: 4766-4774.
- Shi Z, Zhou Y, Yang Z. Approach the contamination of dexamethasone in the effluent water. Guide China Medicine 2012; 10: 319-321.
- Wuthiekanun V, Smith MD, White NJ. Survival of Burkholderia pseudomallei in the absence of nutrients. Trans R Soc Trop Med Hyg 1995; 89: 491.
- Parke JL, Gurian-Sherman D. Diversity of the Burkholderiacepacia complex and implications for risk assessment of biological control strains. Annu Rev Phytopathol 2001; 39: 225- 258.
- Chain PS, Denef VJ, Konstantinidis KT, Vergez LM, Agullo L, Reyes VL, Hauser L, Cordova M, Gomez L, González M, Land M, Lao V, Larimer F, Lipuma JJ, Mahenthiralingam E, Malfatti SA, Marx CJ, Parnell JJ, Ramette A, Richardson P, Seeger M, Smith D, Spilker T, Sul WJ, Tsoi TV, Ulrich LE, Zhulin IB, Tiedje JM. Burkholderiaxernovorans LB400 harbors a multi-replicon, 9.73-Mbp genome shaped for versatility. ProcNatlAcadSci USA 2006; 103: 15280-15287.
- Yi W, Zhibang Y, Lili Z, Zhongquan S, Lianju M, Ziwei T, Renju J. Isolation and identification of dexamethasone sodium phosphate degrading Pseudomonas alcaligenes. J Basic Microbiol 2015; 55: 262-268.
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 1997; 25: 4876-4882.
- Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5:molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods.MolBiolEvol 2011; 28: 2731-2739.
- Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. MolBiolEvol 1987; 4: 406-425.
- Li R, Li Y, Kristiansen K, Wang J. SOAP: short oligonucleotide alignment program. Bioinformatics 2008; 24: 713-714.
- Lagesen K, Hallin P, Rødland EA, Staerfeldt HH, Rognes T. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 2007; 35: 3100-3108.
- Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 1997; 25: 955-964.
- Delcher AL, Bratke KA, Powers EC, Salzberg SL. Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 2007; 23: 673-679.
- Tatusov RL, Natale DA, Garkavtsev IV, Tatusova TA, Shankavaram UT, Rao BS, Kiryutin B, Galperin MY, Fedorova ND, Koonin EV. The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res 2001; 29: 22-28.
- Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M. The KEGG resource for deciphering the genome. Nucleic Acids Res 2004; 32: 277-280.
- Uhía I, Galan B, Morales V, García JL. Initial step in the catabolism of cholesterol by Mycobacterium smegmatis mc2 155. Environ Microbiol 2011; 13: 943-959.
- Uhía I, Galan B, Medrano FJ, Garcia JL. Characterization of the KstR-dependent promoter of the gene for the first step of the cholesterol degradative pathway in Mycobacterium smegmatis. Microbiology 2011; 157: 2670-2680.
- Donova MV, Egorova OV. Microbial steroid transformations: current state and prospects. ApplMicrobiolBiotechnol 2012; 94: 1423-1447.
- Garcia JL, Uhia I, Galan B. Catabolism and biotechnological applications of cholesterol degrading bacteria. MicrobBiotechnol 2012; 5: 679-699.
- Malaviya A, Gomes J. Androstenedione production by biotransformation of phytosterols. BioresourTechnol 2008; 99: 6725-6737.
- Sripalakit P, Wichai U, Saraphanchotiwitthaya A. Biotransformation of various natural sterols to androstenones by Mycobacterium, sp. and some steroid-converting microbial strains. J MolCatal B Enzymatic 2006; 41: 49-54.
- Vasilevskaya AV, Yantsevich AV, Sergeev GV, Lemish AP, Usanov SA, Gilep AA. Identification of Mycobacterium tuberculosis, enzyme involved in vitamin D and 7-dehydrocholesterol metabolism. J Steroid BiochemMolBiol 2016; 30151-30160.